05/15
2020
Improving the sensitivity of the HybriDetect 2T
How to use block oligos for better results
Detecting various PCR amplificates with HybriDetect 2T
The amplification of nucleic acids has become an important tool for various areas such as medical diagnostics, food safety and research over the past decades. A quick and easy way to get a visible result of the amplification is to evaluate the amplification product using the HybriDetect 2T strip, a lateral flow evaluation strip.
The most common method for nucleic acid amplification is the polymerase chain reaction (PCR). The operating principle of the HybriDetect 2T strip is described in figure 1.
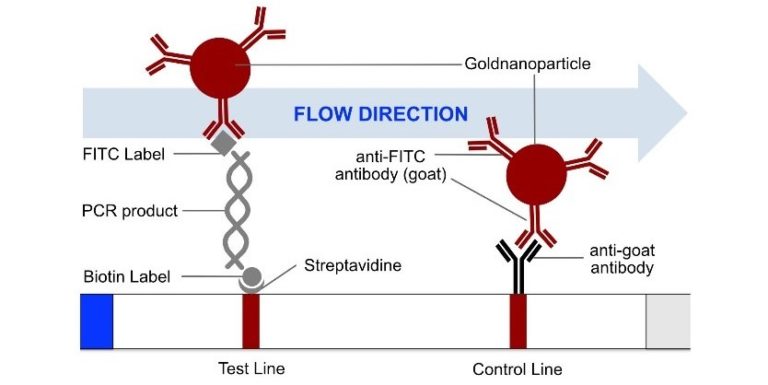
Fig. 1: HybriDetect 2T principle with PCR sample. labeled PCR amplificate can bind to the test line on the strip using biotin label. Bound sample gets visible through gold nanoparticles, which bind the amplificate via FITC
More sensitivity with higher Volume?
Although PCR amplificates are already good to evaluate with the LF strip, there still is potential for an even more sensitive detection. Currently, the optimal application volume for the LF evaluation is only 2 μl out of the total of 25 μl PCR product. If a larger volume is applied, the intensity of the signal does not increase, as one would presume, but decrease (see fig. 2). This means that only a small fraction of the amplified fragment is actually used for detection. Therefore the evaluation could be much more sensitive, if it would be possible to use the entire amount of PCR product.
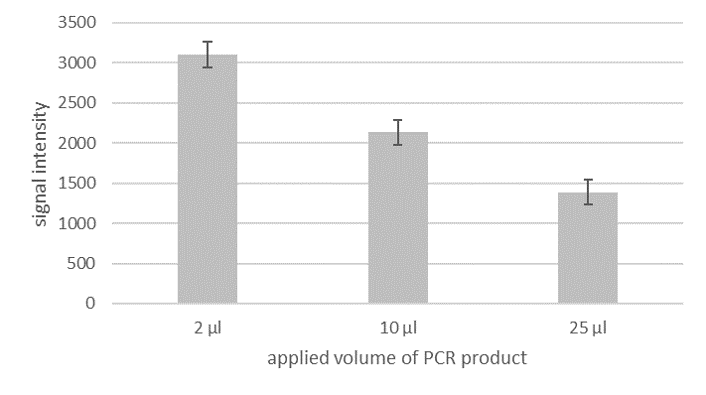
Fig.2: Applying larger Volume leads to a loss of sensitivity. Signal intensity of HybriDetect 2T with 2 µl, 10 µl and 25 µl of PCR product.
Free primers causing loss of sensitivity
The loss of sensitivity mentioned above can be explained as follows: To receive as much amplificate of the original DNA sequence as possible, the marked primers are added in excess to the PCR mix. Hence, some free primers, which are not used in the reaction, remain in the PCR product. Due to their markings, these free primers are able to bind to the binding sites of the lateral flow strip (and the binding site of the chromophoric gold conjugate) in competition to the double stranded PCR amplificate as shown in figure 3. Regarding the FITC marked primer, this means that the primer is able to bind onto the anti-FITC antibodies loaded with gold particles in the same way as the PCR product that is to be detected, so that the resulting signal of the PCR product is attenuated.
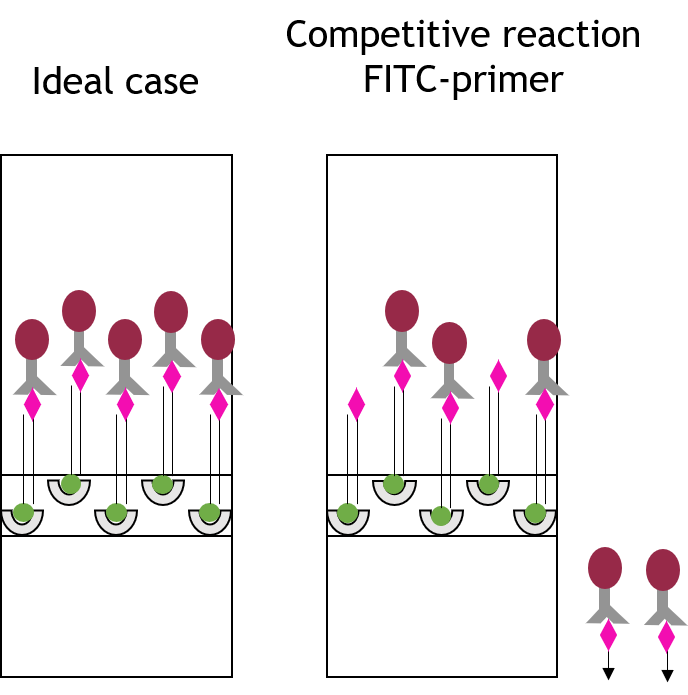
Fig.3: Loss of sensitivity due to competetive reaction. Comparison of the signal intensities of the ideal case evaluation and the real evaluation, containing the competetive reaction between free FITC labeled primers and PCR product.
Designing block oligos to overcome the loss of sensitivity
An idea to compensate the attenuation of the signal and thus loss of sensitivity caused by the free primers, is reflected in the design of extended and modified antisense sequences of these primers, so-called block oligos. The block oligos consist of the antisense sequence of the corresponding, a few spacer nucleotides, a hairpin structure and additional nucleotides serving as overhang (see fig. 4).
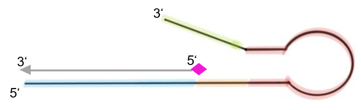
Fig.4: Structure of block oligos to overcome loss of sensitivity. Antisense sequence (blue) bind to free detection primer. Spacer (orange), hairpin (red) and overhang (green) ensure that FITC label (pink) of the detection primer (gray) is shielded.
The idea behind this design is that when these block oligos are added to the PCR mix, after the PCR but prior to the evaluation, they bind to the remaining free primers because of the antisense sequence and the remaining structural elements of the block oligo mask the FITC label of the free primers so that the free primers bound to the block oligos can no longer be bound to the gold conjugate.tag verwenden
Effect of the designed block oligos
To test the effect of the designed block oligos, a test system was developed in which the signal intensity of the block oligo sample was compared with the maximum possible signal intensity (artificial sample without free FITC labeled primers) and with the currently achievable signal intensity. By adding the block oligos to the PCR product and evaluating 25 μl instead of only 2 μl amplificate, a tripling of the signal intensity and thus a tripling of the sensitivity could be reached, as shown in figure 5.
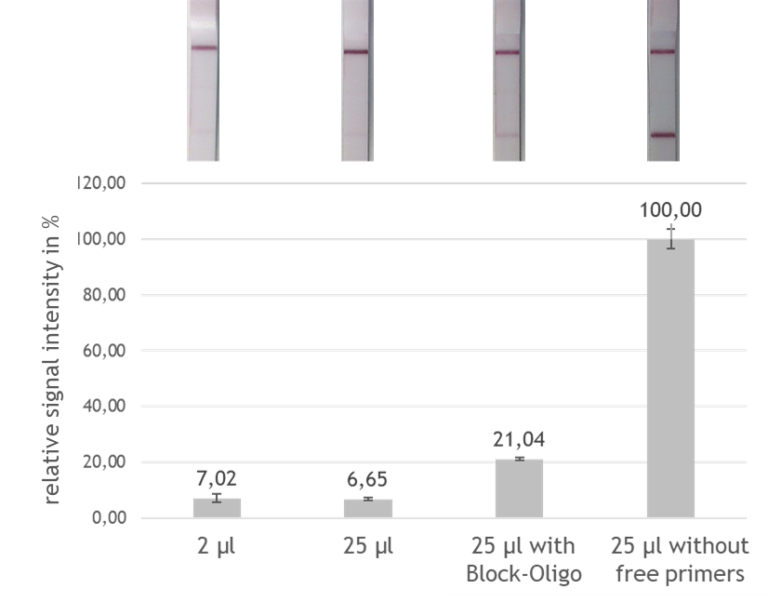
Fig 5.: Effect of the block oligos. Relative signal intensity of the various evaluation variants.
Summary
Regarding the evaluation of PCR amplicons, it was found that the free detection primers, the FITC labeled primers in particular, are responsible for a loss of sensitivity in the evaluation. These free primers can bind to the gold conjugate of the evaluation strips in competition to the PCR amplificate. An improvement for this problem could be achieved with the help of so-called block-oligos, which are added to the PCR product together with the running buffer and whose structure allows them to bind to the free primers and shield their FITC labeling.
Author
Dr. Jacqueline Hoffmann
QA — RA
jhoffmann@milenia-biotec.de
+49 (641) 948883 – 0
Related Products
HybriDetect — Universal Lateral Flow Assay Kit
The HybriDetect is a simple and quick tool to develop your own rapid test. Various molecules can be detected: Proteins, Antibodies, Genetic amplicons
Related Articles

Diagnostic CRISPR Tools and Techniques
Lateral Flow Assays using Genome Editing Tools

HybriDetect and Temperature Effects
Using higher Temperatures: Part of the NAFLIA Toolbox